Most of us have heard that ATP is the source of energy for the cell, along with various explanations of where that energy comes from, but unless someone has studied the biochemistry and biochemical thermodynamics of the cell it's hard to put everything you can read about this into a useful picture. Therefore I'm going to develop an analogy to help explain the various mechanisms used to move energy around the cell.
The Electric Wiring Analogy
Just as a house is wired for 110V electricity, the cell is "wired" for energy. The standard outlet socket, equivalent to the normal plug in your wall, we'll call the "A-socket". This is ATP, as normally used. Just as the wiring is everywhere in the house, the A-socket is available everywhere in the cell (with the exception of a few types of endosome). Now imagine that, when the electrical distribution system was just beginning, different countries had mandated different sockets for outlet plugs, and before everything was standardized so many different manufacturers had created so many useful things using these different socket types that everybody simply wired their houses with all types of socket, so they could use any power appliance (or whatever) they bought without worrying about the plug.
Although this isn't the actual reason exactly (see below), the cell is actually wired with a bunch of different socket types besides the A-socket. To start with, there's the G-, C-, and U-socket. In addition, there are what I'll call "prime" sockets: The A'-, G'-, C-', and T'-sockets. (For historical reasons, there are no T- or U'-sockets.) They all run at the same "voltage", that is the same energy level, however some rooms have all the sockets attached to the same high-capacity wiring while others may have the "funny" sockets attached via thin wires to a single connection to the high-capacity A-socket system. In the latter case, too much use of an appliance attached to one of the "funny" sockets, especially one of the prime sockets, might cause a partial brown-out.
220 Volts
What about the appliances in our house that run on 220V? In fact, there is a very good analogy for this in the splitting of a phyrophosphate group off of ATP rather than the normal phosphate group. The charging of transfer RNA with its appropriate amino acid is an example of this, as is the manufacture of cyclic nucleotide monophosphates. We'll call the equivalent plug for ATP the 2A-socket, while the equivalents of the other will be the 2G-, 2C-, 2U-, 2A'-, 2G'-, 2C'-, and 2T'-sockets.
As it happens, there are some critical appliances in the cell that require not just one but four different types of "220V" sockets, for reasons that break down the analogy somewhat (see below). The system that creates new RNA in genetic transcription (RNA polymerase) requires the 2A-, 2G-, 2C-, and 2U-sockets, while the systems that create new DNA (DNA polymerases) (of which there are several) require the 2A'-, 2G'-, 2C'-, and 2T'-sockets. Thus, these "200V" systems are essential for the most fundamental systems in maintaining life.
Identity of the (Funny) Socket Systems
You may have guessed by now what these other sockets correspond to: the (2)A-, (2)G-, (2)C-, and (2)U-sockets are the ribonucleotide triphosphates, while the (2)A'-, (2)G'-, (2)C'-, and (2)T'-sockets are the deoxyribonucleotide triphosphates.
The reason that the RNA and DNA polymerases require these four types (each) is that they actually create their product out of the nucleotide bases that carry the phosphate chains, along with one of the phosphates. Breaking off the pyrophosphate (i.e. the "220V" plug) provides the energy to create each link in the RNA and DNA chain. As you can see, this process sits entirely outside the "electrical wiring" analogy, since normal appliances (or other power tools) don't consume the wires or outlet plugs they're plugged into. (They're not supposed to, anyway).
If you hadn't guessed what the various socket types corresponded to, don't be disappointed, it's actually an extremely surprising fact for most people that the cell's most powerful energy systems are so fundamentally interlinked with the replication and transcription systems.
Thus, the actual reason that all these "funny" socket systems exist is the need for them in replication and transcription. However, since they're there, a mutation that created a useful enzyme using one of them will likely survive and prosper. It's important to realize that in evolution, there is no "parsimony" of mechanisms. If a mutation forms a mechanism, or ties to an existing one, it gets selected just like one that uses the most common system.
Low-Voltage Systems
Just as many electronic appliances you can buy today use lower voltages, and require some sort of adapter to use house wiring, the cell has its own low-voltage systems, based on membranes: the p-socket system (the lower case "p-" because it stands for "proton", not "Phosphate"), the S-socket system, as well as other special-purpose systems we won't give names to (or discuss further here). All of these membrane-based systems involve the movement of ions (electrically charged molecules) across membranes.
The p-Socket System
The p-socket system is based on proton pressureA1, a combination of relative proton concentration (pH), and voltage across the membrane. Since pH is also a measure of acidity (or rather, acidity is the effect of proton concentration, so pH refers to both), you can see that only one side of a membrane can be at a desired acidity, while the other has to be different by an amount needed to support proton pressure. Fortunately, voltage across the membrane also plays a part, so the energy available from this system is not completely limited by what differences in pH the cell can support on each side of a particular membrane.
The p-socket system is extremely important: of the three major sources of energy transmitted by the A-socket system, two actually feed into the p-socket system and go through a "transformer" (see below) to reach the A-socket system.
The S-Socket System
The S-socket system is based on sodium ion pressure, and like the p-socket system involves both concentration and voltage differences across the membrane. In eukaryotes it's usually different membranes from the p-socket system, primarily the plasma (cell) membrane and the endoplasmic reticulum (ER). Prokaryotes, mitochondria, and chloroplastsA2 have both in one membrane, but most of the energy goes through the p-socket system, with the S-socket system primarily maintaining the sodium ion balance. Nevertheless, it's there, and some enzymes have evolved to use it as a power source.
The most important function of the S-socket system, at least for thinking humans, is the plasma membrane of neurons, where it serves to provide the power for both action potentials and the sophisticated calculations performed in the synaptic and dendritic membranes. Of course, the power originally comes from ATP, via a "sodium pump" that moves sodium and potassium across the membrane.
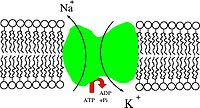
Figure 1: Cartoon of a sodium pump: Na+/K+-ATPase. (From Wiki)

Figure 2: Ribbon diagram of structure of Na+/K+-ATPase. (From Wiki)
Thermodynamics and Kinetics of Cellular Wiring
Let's start with some basics. Energy in the cell comes from chemical reactions. If you're coming at biochemistry as I did, with a background in physics and thermodynamics used for calculations around refrigeration and heat engines (or atmospheric radiation thermodynamics), you're likely to miss, as I did, some of the important aspects of chemical thermodynamics as they apply to biochemistry.
Let's start with the relationship between concentration and energy: If the energy yielded from a specific reaction under certain conditions is x, electron volts (eV), then reducing the concentration of one of the substrates, that is one of the chemicals going into the reaction, will decrease the energy yield by about 0.06eV. (The number is actually 0.05916,1 although I've seen it given as 0.0582 as well; for our purposes 0.06 is good enough, especially since the discrepancy is probably due to different errors in different measurements: there's a lot less precision in "standard" measurements than the precision of the numbers would suggest.1)
Let me say a word about energy measurements, and other units, before I go on to examples: to me, the proper unit of energy for a reaction involving one (or a few) molecule(s) is the electron volt, that is the energy required to move an electron (or other single charge) across a voltage difference of one volt. Chemists normally use KiloJoules/mol, which makes sense for them in measuring reactions in test tubes. But getting a test tube full of living cytoplasm is out of the question, and our theoretical investigations are better served by a unit that applies directly to a single molecule. Similarly, I'm going to measure concentration in molecules per cubic micron (µm-3).A3
The issue with energy and concentration is thermodynamic, it has to do with how entropy interacts with molecules in solution. In addition, various bonds have various amounts of energy, and a high-energy bond will want to hydrolyze (that is, split apart by putting the hydrogen from water on one end and the hydroxide group on the other), much more than a low-energy bond.
How much a bond wants to be hydrolyzed, however, doesn't control how fast it will be. This is the area of kinetics. Some bonds are very slow to hydrolyze on their own, requiring a catalyst to speed them up. The ability of a bond to hydrolyze can be measured by its "half-life", which can be measured in weeks for a bond like that connecting two phosphate groups.
Life is all about kinetics: a specific set of reactions is enabled by enzymes (and ribozymes), while the rest are left with their very long half-lives, effectively unable to happen. (Of course, the living cell is always turning some of its enzymes on and off, so some reactions can only happen at appropriate times.)
ATP and its Relatives: the A-Socket System
ATP stands for Adenosine TriPhosphate, which consists of an organic molecule (adenosine) hooked to three phosphate groups.
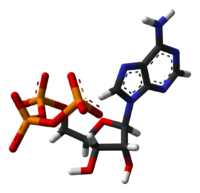
Figure 3: stick model of ATP
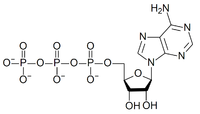
Figure 4: atomic structure of ATP
The "P-O-P" bond is a high-energy bond, which means that when it is hydrolyzed, that is when a water molecule is added to create two molecules ("P-O-H H-O-P") energy is gained. The process ends up creating adenosine diphosphate and inorganic phosphate (Pi). The process of splitting the last phosphate group is the A-socket. Under typical conditions the energy of this reaction will be about 0.6eV. The 2A-socket gives about twice that much by splitting off a pyrophosphate, creating adenosine monophosphate (AMP).

Figure 5: Ball and Stick model of pyrophosphate. (From Wiki.)

Figure 6: chemical formula of AMP. (From Wiki.)
The reason is two very important enzymes, or rather enzyme families: Adenylate Kinase and inorganic pyrophosphatase.
Inorganic pyrophosphatase freely splits the pyrophosphate into two phosphate groups, so that they try to reach equilibrium. Because of the energy in the P-O-P bond, that would work out to less than a millionth of a molecule (ion) per cubic micron (10-6 µm-3). Usually there's a lot more than this, as the pyrophosphate ions created through use of the 2A-socket (and all the other 2- sockets) wander around looking for a molecule of inorganic phosphatase. This means the energy from the 2A-socket (and the others) will actually be somewhat lower than double, the A-socket, but not by much.
The reason for that is the adenylate kinases, (AK's) which convert both ways between ATP+AMP<==>ADP+ADP, with an equilibrium ratio of about [AMP]=~1.12x[ADP]2/[ATP]. (I'm using the brackets to indicate concentration.) As I mentioned, there's a whole family of AK's, AK1 through AK7, all of which perform this function, in different ways.3 They are located in different compartments of the cell, and have slightly different kinetics, and sometimes different substrates: AK3, inside the mitochondria, works with Guanosine Triphosphate (GTP: the G-socket) rather than ATP, allowing the GTP created by the Krebs Cycle to be converted to ATP before it leaves the mitochondrion.
AK's of all types are very sensitive to a variety of conditions, modifying their effectiveness as catalysts by several orders of magnitude. This, combined with the number of enzymes that respond to AMP concentration, puts the AK's at the center of a large, complex web of cellular control.3, 4
It's as though your house had a big computer watching your energy use in every room, anticipating changes in usage, and adding generating capacity and improving the wiring as needed. Both short-term and long term changes are mediated through AMP concentration, which results from the combination of energy usage (via ATP) and the sensitivity of the AK's to various conditions within the cell. Even the cell's ability to grow and change its shape is dependent on modifications to ATP levels by AK's.5
Three Relevant papers
Three recent papers address this subject, and I want to touch on them briefly before going on to the low-voltage systems.3, 4, 5
Dzeja, P., & Terzic, A. (2009). Adenylate Kinase and AMP Signaling Networks: Metabolic Monitoring, Signal Communication and Body Energy Sensing International Journal of Molecular Sciences, 10 (4), 1729-1772 DOI: 10.3390/ijms10041729
Saks, V., Monge, C., & Guzun, R. (2009). Philosophical Basis and Some Historical Aspects of Systems Biology: From Hegel to Noble - Applications for Bioenergetic Research International Journal of Molecular Sciences, 10 (3), 1161-1192 DOI: 10.3390/ijms10031161
van Horssen, R., Janssen, E., Peters, W., van de Pasch, L., Lindert, M., van Dommelen, M., Linssen, P., Hagen, T., Fransen, J., & Wieringa, B. (2008). Modulation of Cell Motility by Spatial Repositioning of Enzymatic ATP/ADP Exchange Capacity Journal of Biological Chemistry, 284 (3), 1620-1627 DOI: 10.1074/jbc.M806974200
The first, Adenylate Kinase and AMP Signaling Networks: Metabolic Monitoring, Signal Communication and Body Energy Sensing, describes the enormous number of control reactions that the AK's are involved in. Since they modulate the concentrations of AMP in association with energy usage, specifically local ATP usage, this ties the cell's primary energy wiring directly into a very complex control system. Everything the cell does takes energy, and by modifying their kinetics the AK's can anticipate, modify, or suppress the response of many other systems to local energy usage. This signaling system extends outside the cell as well, extra-cellular ATP and AMP are part of the control system.3
The second paper, Philosophical Basis and Some Historical Aspects of Systems Biology: From Hegel to Noble - Applications for Bioenergetic Research, takes a more philosophical look at the implications of recent discoveries regarding the A-socket (and 2A-socket) systems and their integration into the cells computing systems. It stresses "new perspectives for studies of the integrated processes of energy metabolism in different cells. These integrated systems acquire new, system-level properties due to interaction of cellular components, such as metabolic compartmentation, channeling and functional coupling mechanisms, which are central for regulation of the energy fluxes." This is very important: the older, reductionist, approach to research needs to be joined by a more holistic, approach: "Systems Biology applies methods inspired by cybernetics, network analysis, and non-equilibrium dynamics of open systems."
The third paper, Modulation of Cell Motility by Spatial Repositioning of Enzymatic ATP/ADP Exchange Capacity, investigates the specific interaction between local concentrations of ATP and the cell's ability to "crawl" through active restructuring of the actin cytoskeleton. By developing a way to locally control the concentration of one of the adenylate kinases, AK1, they were able to modify the rate of motility and spreading by the cells. This demonstrates that "[l]ocal ATP/ADP exchange enhances cell spreading [and] motility." More generally, it demonstrates how the AK's participate in a cell-wide system of calculation and control that adjusts the local energy conditions to fit immediate needs.
The rest of the Tri-Phosphates
The other tri-phosphate sockets, the G-, C-, U-, A'-, G'-, C'-, and T'-sockets, and the 2G-, 2C-, 2U-, 2A'-, 2G'-, 2C'-, and 2T'-sockets, are all maintained in energy states similar to the A- and 2A-sockets by enzymes that convert among them. There are control systems that maintain needed levels of these enzymes, but I'm not going to discuss them, since, to the extent we know about them, they're pretty similar for our purposes.
Where the Energy Comes From
There are three primary sources of energy for the cell, and, as mentioned above, two of them involve the p-socket system. Let's start with the one that doesn't, glycolysis. I'm not going to discuss the details here, but in general it takes a sugar molecule and breaks it up and rearranges it to create to pyruvate molecules. The molecular rearrangement produces enough energy to tack extra phosphate groups onto two ADP's, creating two ATP's. (It also creates two molecules of NADH, which can enter the mitochondria and the electron transport chain, but they don't have to, the extra electrons can be dumped onto the pyruvates to create lactates, a fully anaerobic process.)
The second and third processes that create ATP involve the electron transport chain. In this mechanism, high-energy electrons have been attached to a molecule called nicotinamide adenine dinucleotide (NAD) to create NADH. (When it gives up its two electrons, the result is the charged NAD+ and a hydrogen ion: H+.) The electrons go through a series of steps, moving from one membrane-bound enzyme to another via a series of progressively lower-energy electron acceptors, first an oil-soluble quinone in the middle of the membrane, then a free-floaing (water soluble) cytochrome, finally four of them landing on an oxygen molecule. Each step in the chain is performed by a large trans-membrane enzyme complex which uses the energy from the electron to move a number of protons across the membrane, adding energy to the p-socket system.
The transformer that converts between the p-socket system and the A-socket system is called ATP Synthase.

Figure 7: Cartoon of ATP synthase. (From the ATP synthase web page).
ATP Synthase actually is a motor, the stalk that sticks up into the lumen of the mitochondrion (or chloroplast) rotates relative to the part that's buried in the membrane. Each rotation creates three ATP molecules, at the cost of 9-12 protons passing through the membrane. This motor is fully reversible: if the concentration of ATP gets high enough relative to the proton pressure, it can run in reverse and "pump up" the p-socket system.
But Where do the High-Energy Electrons Come From?
These can be created by oxidation of a C-H (Carbon-hydrogen) or C-C bond, in the mitochondria, or they can be created by chlorophyll as part of photosynthesis. But that's a subject for another post.
P.S. See also Leonardo Da Vinci and the F0-F1 ATPase for a simulated movie of ATP synthase.
P.P.S. (6/01/09) If you liked this post, you can vote for it in the 3 Quarks Daily 2009 Science Prize. Or you can check out the opposition here, first. (Lot's of good stuff there.)
Appendices I've taken thoughts that were too rambling out of line here.
A1. "proton pressure":
These aren't the same as the "protons" studied by physicists. Rather, the word is used for a hydrogen ion (H+). However, H+ doesn't actually occur in aqueous environments, instead there are hydronium (H3O+) ions. However, movement is actually faster than a molecule that size, or even if they were single protons. What actually happens is that one of the hydrogens in the hydronium "unbonds" itself from its oxygen and bonds to a new one, so that the charge, and the effective hydronium, has moved over without anything heavier than an electron actually moving.
A2. "chloroplasts":
It's the fad today to call these plastids, in honor of the fact that there are some types without chlorophyll, however I prefer to discuss chloroplasts, as that's almost certainly how they originated.
A3. "[...] is the electron volt [...] molecules per cubic micron (µm-3)."
Conversion Factors:
- 1 eV = 96.4539 KJoules/mol
- 1 molar = ~6.022 x108 µm-3
Typical Concentrations:
- ATP: 6-54 x105 µm-3(9)
- ADP: 2.5-12 x105 µm-3(9) but these numbers don't allow for the fact that most ADP is bound to F-actin in muscles and maybe many other types of cell.7 Free concentrations have been estimated at 9-90 x103 µm-3 (8)
- AMP: 1-3 x105 µm-3(9) although another source shows the very broad range of 54-15000 µm-3 (.8)
- Pi: 1.5-15 x105 µm-3(9) although this can reach levels of 1.2 x107 µm-3 during extreme exercise.8
References and Links:
1. Thermodynamics of Natural Systems by G. M. Anderson
2. The Neuron Cell and Molecular Biology by Irwin B. Levitan and Leonard K. Kaczmarek
3. Adenylate Kinase and AMP Signaling Networks: Metabolic Monitoring, Signal Communication and Body Energy Sensing
4. Philosophical Basis and Some Historical Aspects of Systems Biology: From Hegel to Noble - Applications for Bioenergetic Research
5. Modulation of Cell Motility by Spatial Repositioning of Enzymatic ATP/ADP Exchange Capacity (Preliminary PDF)
6. Oxidative ATP synthesis in skeletal muscle is controlled by substrate feedback
7. Skeletal Muscle Fatigue: Cellular Mechanisms
8. Protecting the cellular energy state during contractions: role of AMP deaminase
9. The Contents of Adenine Nucleotides, Phosphagens and some Glycolytic Intermediates in Resting Muscles from Vertebrates and Invertebrates
10. Roles of the creatine kinase system and myoglobin in maintaining energetic state in the working heart
11. Interrelations of ATP synthesis and proton handling in ischaemically exercising human forearm muscle studied by 31P magnetic resonance spectroscopy
12. Cytosolic phosphorylation potential

No comments:
Post a Comment