One thing I didn't discuss, though, was memory: the ability to store information regarding a specific past state for use in future calculations. While there are several types of memory available to the cell, the one I'm going to talk about here is DNA Methylation, which is a way the cell has of modifying its DNA to carry information into future generations. Unlike in the series on cell smarts, I'm going to was a little more speculative, which is really the purpose of my blogging. Of course, such speculation has to be properly grounded in good scientific research. Nevertheless, this subject can be communicated mostly with references to Wikipedia and more illustrative websites.
The basis of DNA methylation is that one of the bases, usually cytosine, has a methyl group added to its #5 carbon.

Figure 1: DNA Methylation Reaction Catalyzed by DNA Methyltransferase (DNMT), an enzyme that can perform this function. Click on image to see full illustration. (From the website of Keith D. Robertson, Ph.D. Assistant Professor.)
Just adding a methyl group to one base isn't going to create inheritable memory, but it happens there is another mechanism that works during replication to transfer the information to the next generation.
Most DNA methylation takes place at a CpG or CpNpG sequence on one strand. Here the lower case "p" stands for phosphate (linking two nucleic acids) and the upper "N" stands for any base. Now, since the other strand runs in the opposite direction, you can see that there will be an identical CpG or CpNpG sequence on that strand as well, since C (Cytosine) and G (Guanine) hook together in double stranded DNA.
There are several enzymes, expressed at appropriate times and places, which will take a semi-methylated CpG or CpNpG sequence and finish the job on the unmethylated cytosine.

Figure 2. Ball model of DNA with methyl groups in red. Note how the groups on cyctosines on opposite strands are close to each other. (From the website of Keith D. Robertson, Ph.D. Assistant Professor.)
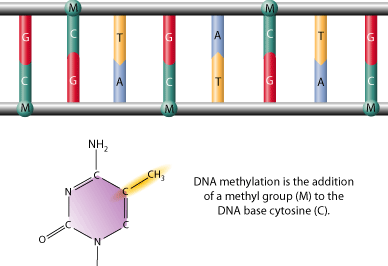 Figure 3: An untwisted double strand of DNA showing several occurrences of CpNpG sequences methylated on both strands. (From HUMAN CLONING: SCIENCE FICTION OR REALITY? by Grace Yim.)
Figure 3: An untwisted double strand of DNA showing several occurrences of CpNpG sequences methylated on both strands. (From HUMAN CLONING: SCIENCE FICTION OR REALITY? by Grace Yim.)DNA methylation is one common system of epigenetic memory, interacting with histone modification and something called RNA interference to modify how the DNA is transcribed going forward in future cell generations.

Figure 4. DNA Methylation and Histone modification. Note that there's one example of CpG methylation, and one example of CpNpG methylation. (From Molecular Development - Epigenetics by Dr Mark Hill.)
The known operation of DNA methylation in epigenetic inheritance (inheritance of acquired characteristics) involves three items too important to leave out, even though they're tangential to what I'm going to discuss. The first is "epimutation", in which a previously stable pattern of methylation (or other epigenetic marking) changes without any change to the base sequence of the DNA. An important aspect of this is that there may be enzymes that control the methylation sequences under certain circumstances, so epimutations could work together with specific cellular mechanisms to provide innovation for Darwinian evolution.
The second item involves regular mutation, in this case the specific change of methylated cytosine to thymine through loss of the amine group. According to The rate of hydrolytic deamination of 5-methylcytosine in double-stranded DNA by Jiang-Cheng Shen, William M.Rideout Ill and Peter A.Jones,
The modified base, 5-methylcytosine, constitutes approximately 1% of human DNA, but sites containing 5-methylcytosine account for at least 30% of all germline and somatic point mutations.
This isn't just because it mutates easier, although according to Shen et al. "The rate constants for spontaneous hydrolytic deamination of 5-methylcytosine and cytosine in double-stranded DNA at 370C were 5.8 x10-13 s-1 and 2.6x10-13 S-1, respectively", but also because the change creates a valid base, which cannot be recognized by the error-correcting machinery of the cell. Conversly, the deamination of regular cytosine creates a "weird" base which the error-correcting machinery will respond to.
The third item involves speciation. The processes by which one population of a species splits into two populations whose interbreeding rate is small enough that they can constitute two species appear to be many, but one of them seems to involve changes in epigenetic marking that make members of the two populations incompatible. This could happen either through mutations to the actual system of enzymes that control methylation, or through the silencing of different defective genes so that any cross-breed ends up with a bunch of unsilenced defective genes and doesn't survive.
A good book about speciation in general is Speciation By Jerry A. Coyne, H. Allen Orr, while epigenetic memory is discussed by two of the experts in Evolution in four dimensions by Eva Jablonka, Marion J. Lamb, Anna Zeligowski.
Now that I've provided a very sketchy description of DNA methylation with links to more, I'll get into the speculation.
In principle, methylation of DNA provides an independent system of memory for the cell, which is not only carried from one cell generation to the next, but can be modified by enzymes within the cell, and sampled by other enzymes so that what's remembered can be input to cellular calculations.
Imagine, then, an enzyme that could recognize a specific sequence of 5-10 bases, and sample a CpG or CpNpG group at a specified point near those bases to see if it's methylated. At the same time, another enzyme recognizes the same specific sequence of 5-10 bases, and methylates the CpG or CpNpG group. A third demethylates it.
If the enzymes were used in appropriate quantities, this one sequence could constitute a binary "switch" whose value could be set, sampled, and inherited through many generations. Of course, it would be better to have a few dozen, or hundred instances of the specific sequence in question, so that if a few were lost to mutation the information wouldn't be lost.
If only five bases were used, potentially 1024 switches would be available. If 10 bases were used, over a million switches. Assuming 50 instances of each switch, that would add up to 50 million bases, less than 1/10th of a percent of the human genome. With 50 instances, the occasional mutation would be little loss for the switch it had been, and even if its methylation value was opposite from that of all the other switches in its new sequence, it would be a small minority. It would be easy enough for a cell to have a "cleaning cycle" in which the value of each switch was first sampled then reset to the value found, wiping out any epimutations.
A million switches would add up to a million bits of information, 125 kilobytes, a pretty good amount. Of course, it would also require a million different enzymes to recognize a million different sequences. However, a few thousand switches would certainly be feasible. In addition, it's possible that RNA sequences could be used to actually perform the recognition.
There is another, more interesting, point regarding DNA methylation as memory, which involves the huge quantities of so-called "Junk DNA", that is DNA for which no use has yet been found. One of the largest types of this "Junk DNA" is called "Alu repeats" after the restriction endonuclease originally used to find it. (In Genome-wide tracking of unmethylated DNA Alu repeats in normal and cancer cells by Jairo Rodriguez, Laura Vives, Mireia Jordà, Cristina Morales, Mar Muñoz, Elisenda Vendrell, and Miguel A. Peinado, is an analysis of available CpG sites and methylation rates in healthy and cancerous mammalian cells.)
The point of this is that we find the same sequence, or very close relatives to it, repeated millions of times. How could this constitute a switch? It probably couldn't, or at least not a very cost-effective one. However, it could constitute a form of analog memory. Consider that there might be a few dozen "starting sequences" that act like promoters for gene transcription. However, instead of transcribing the DNA, the enzyme group (a different one than for transcription) starts at the promoter and runs down the DNA checking each of a particular "switch's" CpG group until it finds one that's unmethylated. The farther it runs down the DNA, the more of some other sort of activity it provides, allowing a roughly analog value to be sampled by the enzyme system.
Thus, analog memory. But what for? Well, there are many areas in the body where a cell has to respond differently according to its location within a specific sheet of cells, especially in the cerebral cortex. It's possible that it finds its position dynamically, according to diffusion from cells at the edge of the sheet. However, another possibility is that it determines its location once, codes it into DNA methylated memory, and thereafter uses it as a reference, input to a variety of cellular calculations that require its location, or other analog parameters. Note that one cell could only use a specific string to store one analog value, but there are enough "Alu repeats" in the human genome for dozens, perhaps hundreds, of analog memory stores, and different types of cells could use the same string for different functions, just as they use the same signals (e.g. cAMP) for different functions.
This might explain why the human genome has so much "junk DNA": it isn't junk but the raw material for "long term memory" in creating and maintaining the brain.
P.S. So why, if all of this stuff is there, hasn't anybody found it? Well, I'm not saying it is there, but that it might be. And as demonstrated in The Genetic Signatures of Noncoding RNAs, by John S. Mattick, "Until recently, most mapping projects have focused on protein-coding sequences, and the limited number of identified regulatory mutations have been interpreted as affecting conventional cis-acting promoter and enhancer elements, although these regions are often themselves transcribed." This means that about the only thing discovered this far involves protein coding, and even regarding regulation, "the limited number of identified regulatory mutations have been interpreted as affecting conventional cis-acting promoter and enhancer elements, although these regions are often themselves transcribed."
In other words, nobody's found it because nobody's looked for it.

I completely agree with you.Great site. A lot of useful information here. I’m sending it to some friends!
ReplyDeleteA behavioral memory's lifetime represents multiple molecular lifetimes, suggesting the necessity for a self-perpetuating signal. One candidate is DNA methylation, a transcriptional repression mechanism that maintains cellular memory throughout development and preserve long-lasting memories. Thanks for sharing this unique information.
ReplyDelete