(A note about footnotes: I've numbered the links at the bottom of the page, and provided numbered annotations in the text. However, since this work isn't intended to be submitted for "peer review" (although any "peer" who wants to review it is more than welcome), most of the footnotes are called out at the section level rather than to support detailed claims. I've also included several links that aren't called out in the text.)
We saw that enzyme activation can take place among a suite of dissolved enzymes, where the changes in concentration are assumed to be homogeneous. (I didn't actually say this, but it was implicit.) But what about when one or more of the participants in the network is localized: perhaps it's bound to a membrane in one spot, or the eyespot of a euglena.
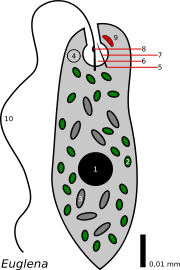
Figure 1: Euglena. The eyespot is #9.
How are such calculations localized? Can we use localization to get more than one calculation at a time out of the same set of enzymes produced by one nucleus?
Methods of Localization
The most important way of localizing calculations is through diffusion.15, 16, 17, 18, 22, 23 When phosphorylization is taking place at one location, the results will diffuse away, towards wherever the dephosphorylation is taking place. The typical scale for diffusion is a micron or so: about the size of a typical bacterium, but 1/10th or less of the scale for a typical eukaryote cell. Some of the larger cells, such as neurons and muscle cells, are so large that diffusion would make it almost impossible for such signals to cross the cell in any useful time. Of course, it also depends on the needed speed. In the neural synapse, response times measured in microseconds are needed, and distances (in, e.g. the PostSynapticPad) are measured in nanometers (the scale of 10-20 atoms in a molecule).
The fastest and most efficient calculations will take place when all the participants in the calculation are localized in one tiny spot. There are trade-offs, however, in size, speed, and accuracy. Try to get too fast, in too small a volume (or membrane-bound area) and the intrinsic fluctuations from individual molecular reactions will introduce too much noise. In the other direction, too much distance will not only reduce the speed, but introduce new fluctuations from random variation in diffusion.
Another way to localize the reactions is by binding one or more of the participants to a membrane.36, 40, 41 Membranes generally act as 2-dimensional liquids, so that means membrane-bound enzymes are, at least, located on one boundary of a diffusible volume.
There are mechanisms to prevent membrane-bound molecules from passing a certain line as well. These won't be much use in a flat membrane, but when it's confined into pouch-shaped regions, the neck of the pouch can be cut off from the rest of the membrane. One mechanism that is useful on a (relatively) flat membrane is rafting, where a number of molecules deeply inset into the membrane (or that pass all the way through it) are hooked together at one location.
Membranes themselves can act as boundaries, enclosing a bunch of dissolved enzymes in a vesicle or endosome.3, 4, 38, 40 These are small globular sections of membrane, typically separated from the larger membranes of the cell. In addition, several types of classic organelles, such as lysosomes and peroxysomes, are small enough to localize "smart" reactions in their lumens. All of these membrane-enclosed organelles can also localize reactions of enzymes bound to their outer surfaces.
Mitochondria, the "power houses" of the cell, are much more than that.34, 35, 38, 39 They are bounded by a double membrane (unlike the organelles mentioned above, or the endoplasmic reticulum), and may localize reactions to the interior, the intermembrane area, or the exterior (for membrane-bound enzymes).

Figure 2: Some of the signaling processes that involve mitochondria. From ref. 35 Figure 1.
There are larger structures bounded by membranes that can also localize computing reactions, such as the endoplasmic reticulum (ER), and the sarcoplasmic reticulum (SR), which is a special type of ER in muscle that serves primarily to contain calcium. These membranes can follow close under the plasma membrane, so that signals (from outside) can cross the short space and cause reactions in membrane-bound receptors on the ER/SR.6, 42
The various parts of the Golgi Apparatus can also localize reactions to their lumens or external surfaces. While the reactions contained in each Cisterna are probably focused on construction activity, at least some of those on the external surface of the membrane need to be involved in guiding the building and fusing of vesicles involved in transporting intermediate products.
The nucleus is a major organelle in the cell (if not always the largest), and its double-membrane envelope provides a number of opportunities for localization.4 The outer membrane is contiguous with the membrane of the ER, however the tube-shaped connections may well have barriers to diffusion between the lumens, as well as along either face of the membrane (for membrane-bound enzymes). The inner membrane appears to have occasional intercourse with outer, as well as some connection through the nuclear pores, however both those connections may be highly filtered, with barriers to random diffusion (ie specific enzymes may be blocked from passing).

Figure 3: Nuclear pore from page linked to in text
The plasma membrane (also called the cell membrane), which is the outer boundary for the cell, can also serve to localize reactions.4, 5, 21, 32, 33 Parts of it can be bounded, as happens in epithelial cells, where the plasma membrane on each side of the tight junction is isolated from the other. In addition, there are intrusions of the plasma membrane into the cytoplasm, called Caveolae, that can isolate both membrane-bound enzymes and free-floating enzymes in the enclosed volume (even though it may be partly congruent with the outside). This may be especially useful in responding to external gradients or localized stimuli, since receptors on the outer surface (localized to the Caveola) can communicate with localized membrane-bound enzymes on the internal surface, participating in a local reaction of any arbitrary complexity (in principle).
In the muscles of mammals, there are organelles called T-tubules which are deep, tube-shaped incursions of plasma membrane into the cytoplasm. These have been observed to have localized processes, including manipulation of the electrical field.

Figure 4: T-tubule and SR in localized reactions. From ref. 32 figure 1.
An organ found in vertebrates called the "primary cilium", which appears to be descended from an earlier flagellum, is sort of like a T-tubule pointing outwards, although it appears to have more sophisticated mechanisms for localizing proteins.7, 8, 9, 10, 11 This organ could detect extra-cellular gradients by using separate calculations at the tip and base. In addition, it could communicate spacial information to the cell as a whole by means of releasing varied concentrations of several enzymes of messenger molecules (see below) at once, corresponding to such scalar values as distance, angle, etc.
Another form of extrusion of the plasma membrane is the dendritic spine of many (most) types of neuron. This organelle (if that's a correct name) can isolate concentrations of cytoplasmic enzymes or other molecules, as well isolating membrane-bound (and -spanning) enzymes on either face. (Membrane faces are sometimes called "leaflets".)
Dendritic spines are very important for understanding cellular intelligence, since they not only can localize enough calculating ability to perform complex analog calculations, but these calculations are intimately involved in the interactions of neurons that produce, among other things, our own sort of intelligence. We'll be seeing more of these spines.
Cellular Scaffolding12
We won't go into the details of the cells internal skeleton, but we should note that there are several ways in which it can be used to localize reactions, or entire calculation units.
One possibility is that a number of molecules of an enzyme, or several enzymes participating in a reaction/calculation, can be simply fixed to one point on the cellular skeleton. They can also be fixed to some other structure that is connected to the skeleton.
Alternatively, they can be attached to some mobile element, such as a mobile vesicle, or directly to one of the cellular motors that drag various things around the cell.
Methods of Signaling (Beyond phosphoactivation)
Once we've begun to localize our calculating reactions, we have to consider the difference between simple calculating elements, and signals. A signal is usually something that carries information from one place to another. Most of them arise at a specific point, either on the outside of the cell, or at some internal organelle or other object. At this point, the speed with which that signal propagates helps to determine how localized the reaction is.
Since phosphoactivating enzymes tend to be pretty large, with slow diffusion rates, the cell has evolved a number of signals with faster propagation.3, 35 First, we'll consider small diffusible things: molecules and ions. (I'm not going to even try to make a complete list.) Cyclic nucleotide monophosphates (cNMP's) are messenger molecules created from nucleoside triphosphates by removing a pyrophosphate group and hooking the remaining phosphate to a second carbon in the ribose. Important cNMP's are cyclic adenosine monophosphate (cAMP) and cyclic guanosine monophosphate (cGMP).
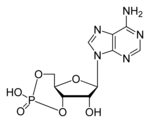

Figure 5: Atomic structure and ball model of cAMP.

Figure 6: Atomic structure of cGMP.
Creation of cNMP's is expensive in energy, costing two ATP (to ADP) reactions for each cNMP. (Separating a pyrophosphate group is converted to separating two phosphate groups from two ATP's via other enzymes in the cell. This is covered in any biochemistry textbook, but I haven't found a good online link.) However, this expense provides an advantage, since the reaction can be driven to extremely high concentrations, driving rapid diffusion. Similar is true for breaking the 3'-bond to create simple monophosphates. Thus, like phosphoactivation, creation of cNMP's achieves speed at the cost of energy.
Inositol triphosphate (InsP3) is another common messenger molecule, along with diacylglycerol (DAG) usually created with it by hydrolysis of Phosphatidylinositol 4,5-bisphosphate, which is a component of many membranes.



Figure 7: InsP3, DAP, Phosphatidylinositol 4,5-bisphosphate. DAP lives deep in the membrane, as does the part marked blue in Phosphatidylinositol 4,5-bisphosphate.
An advantage of this messenger molecule is that the precursor is membrane bound, which means it can be maintained at fairly high concentrations without creating huge amounts of it. Of the two messenger molecules created, one is free-floating, one membrane bound. This means that responses can be hooked to either, depending on circumstances.
InsP3 and some of its relatives tend to be localized to the cytosolic side of various membranes, helping to stimulate and localize several reactions within the cell.

Figure 8: Phosphoinositides and their metabolizing enzymes in exo-endocytic membrane traffic. (From ref 3 Figure 1. Click on image to see original caption.)
One of the most powerful, and common, messengers is calcium, specifically the calcium +2 ion (Ca++). Calcium is probably the smallest, and fastest diffusing, messenger available to life. The proton (technically the positive hydrogen ion, it isn't the same as the naked proton of physics) is faster, but simply too common at a life-sustaining pH for easy use as a messenger.
Other ions, both positive (cations) and negative (anions) may also be used as messengers, however this is much more rare.
Transmembrane ion Channels
Ion channels are pore-forming proteins that allow ions to cross the membrane (which they normally can't do because they're charged). Some are invariant, but the ones we're interested in are those that can react to the presence of various activated enzymes, signal molecules, or other signals (such as voltage). In order for the ions to cross, there has to be a difference in concentration, voltage, or both across the membrane, which in turn has to be maintained by some source of energy, usually an ATP-driven pump. When these channels open, they allow a current of ions across the membrane, with corresponding changes to membrane voltage and/or ion concentration. Calcium channels are particularly important: a very common reaction involves insP3 being released at the plasma membrane, diffusing to the ER, and reacting with a calcium channel in the ER membrane that allows large amounts of calcium to enter the cytoplasm from the lumen.
Voltage across the membrane is another sort of signal,32 one that moves very fast, although not at the speed of light as many would suggest. Voltage is created by allowing a current of some ion that has a very large concentration gradient across the membrane: ions flow "uphill" electrically producing a current against a growing voltage. Depending on the parameters, the voltage may stabilize somewhere against the concentration difference, or may spike and return after the gate closes.
Voltage signals combine with the capacitance and conductivity of the membrane to create a local spot of high voltage that drops and spreads out. If the current is pulsed (starts then shuts off) it starts as a high, narrow, hill, then becomes wider and lower until it fades out as a very wide low bump. Variations in the inputs and characteristics of the ion channel(s) can produce various time-dependent effects in the voltage at different distances. Since some sort of voltage-sensing ability is needed to respond to the signal, which will have some threshold voltage, the signal cannot be considered "light-speed". Nevertheless, speeds of hundreds of meters/second (equivalent to a few nano-seconds to cross 1 micron) can be achieved.
Another signalling mechanism involves trigger mechanisms. For instance, the phosphorylated version of an enzyme might be able to catalyse its own phosphorylation. Thus, the enzyme exists in a completely unphosphorylated state, until some other signal causes a few molecules to become phosphorylated, after which an expanding wave of phosphorylation propagates throughout the extent of unphosphorylated enzyme. Of course, this "trigger" will have to be reset after use, requiring a system of several other enzymes.
One of the most important trigger mechanisms is used by the axons of neurons, where a combination of sodium and potassium gradients, voltage across the membrane, and voltage-gated ion channels combine to produce the moving action potential. In naked axons it can achieve speeds as high as 5 meters/second, (much higher in special neurons,) while myelinated neurons can get up to 50 meters/second.

Figure 9: ratio of axon diameter and theoretical speed for myelinated and unmyelinated cat neurons.
Looking at the illustration, we could project the speed for an unmyelinated tube 1/10th of a micron in diameter as perhaps half a meter/second, equivalent to 10 microsecond to cross 5 microns, a typical size for a small T-Tubule. In principle, a single cell could send signals within its cytoplasm at this rate, although I don't know of any actual examples.
The Calcium System2, 26, 27, 34, 35, 38, 39
Calcium, as mentioned above, is probably the fastest signaling ion available. The normal concentration in the cytoplasm is very small, compared to the external medium, and that within the ER/SR. This differential is maintained against both the concentration gradient and often a voltage by ion pumps, ultimately powered by ATP. (These pumps may be directly powered, or they may use another ion differential (and voltage) as an energy source.) Sodium is the most frequent. The release of Ca++ into the cytoplasm is achieved by ion channels, usually responding to another messenger such as hormones and neurotransmitters outside the cell, and other signaling molecules within the cell. They can also respond to voltage or calcium itself within the cytoplasm: there is a positive feedback mechanism often used to allow calcium released through the plasma membrane to stimulate a much higher amount of release from the ER/SR. (This is a rather complicated mechanism and I won't try to describe it here.)
The mitochondria play a major role in the calcium system.38 They not only serve to buffer the calcium concentration by quickly absorbing much of it as it's released, but the addition of calcium inside the mitochondrion stimulates a local increase in ATP production. This is very important, because it supports the use of calcium to signal all sorts of different energy-using processes within the same cell. Most of the absorbed calcium is returned directly to the lumen of the ER/SR, without going through the cytoplasm. We should note that the buffering acts together with diffusion to strongly localize calcium signals, often to within a small fraction of a micron.
A very common enzyme involved in responding to calcium signals is calmodulin.2
One important example of the calcium signaling system is in the pre-synaptic terminal (in nerve synapses).28, 29 When the action potential along the axon reaches the pre-synaptic terminal, it triggers a strong release of calcium, which in turn causes vesicles filled with neurotransmitters to fuse with the plasma membrane in the synapse and release neurotransmitters. The movement of vesicles, and recovery of neurotransmitters from outside the cell (the synaptic gap is actually outside both nerve cells), are both energy intensive processes, and the ability of the released calcium to stimulate an immediate increase in ATP production means it doesn't have to wait until the ATP is depleted by those processes.
Interestingly, pre-synaptic vesicles are often "pre-fused" with the membrane, stalled partway through the process of release, until the calcium signal appears. This allows a much faster response to the action potential.
Calcium is also involved in most post-synaptic processes, in a very different way. Here, it may be used to stimulate all sorts of different ion channels, helping to control the electrical response of the local plasma membrane. Again, its ability to "pre-stimulate" ATP production in the mitochondria helps keep things on an even keel.
Dendritic Spines1, 13, 14, 21, 22, 24, 25, 35, 36

Figure 10. Cartoon depicting main types of shapes for dendritic spines. (from Wiki)
Many types of neural synapses have a "spine" on the post-synaptic side, isolating the postsynaptic density from the main part of the dendrite it's located on. The head of the spine is around a femtoliter (a cubic micron) in volume, which makes it close to the ideal volume for isolated analog calculations.
Dendritic spines can concentrate membrane-bound enzymes, either in the head or the neck (or both), as well as in the postsynaptic density, which is the part of the actual synapse located within the (receiving) cell. For instance, enzymes that degrade cAMP and/or cGMP can be so isolated, preventing even strong signals from having a significant effect outside the spine itself. We should note that spines are not needed to isolate calcium signals,21 however they also isolate many other reactions involved in the complex calculation that takes place at every dendrite.

Figure 11. Localization of enzyme PDE10A in a section through rat striatum using immuno-electron microscopy. Black dots represent molecules of enzyme, to which gold particles have been attached, allowing them to show up in the electron micrograph. D=dendrite body in cross-section (note the ER in the middle of the tube), P=pre-synaptic terminal (filled with vesicles), S=spine, PSD=post-synaptic density. Blue arrows point to PDE10A bound to dendritic and spine membranes, red arrows to PDE10A within spines not apparently membrane-bound, yellow arrows to PDE10A bound at the edge of the PSD. Click image to see original figure and caption. From ref. 1 Figure 7.
The post-synaptic density contains scaffolding elements as well as a variety of membrane-bound proteins, all of which can help to isolate enzymes and their reactions. When the pre-synaptic nerve cell fires, the neurotransmitters are released into the synaptic cleft and travel to the membrane over the PSD. Here they react with a variety of receptors, triggering any number of local reactions within the dendritic spine.
The most prominent response is an ion current, which can influence the local voltage in the spine, which voltage flows across the neck and influences the dendritic tube. Depending on the specific structure, which varies among the many types of neurons in the brain and body, even one synapse can cause the receiving neuron to fire in its turn, although it's much more common for several input to be needed. Among the ions that can flow in the currents are sodium, potassium, cloride, and, of course, calcium. Some of these ion currents increase the voltage (driven by a high concentration difference), while others decrease it.
When one or more impulses reach a spine, it can sometimes contract, pushing a pulse of calcium and other messenger molecules into the dendrite proper.24 This can work in addition to diffusion to move signals into the dendrites, which can effect local changes in the behavior of the membrane and its cytoplasm.
This is very important in considering just how smart a neuron can be. While the currents in direct response to incoming nerve pulses are usually localized to the PSD, the result is a voltage change that propagates through the neck of the spine and into the dendrite, often reaching the soma (body) of the nerve cell, where it affects when (and whether) the nerve cell fires. The dendritic membrane acts somewhat like a cable in this process, passing along the electrical effect.
But it is not a "passive cable". There are a number of ion channels in the dendritic membrane, different depending on the nerve cell type and its immediate history. Many of these channels are voltage-driven, so that there is an actual current across the membrane, that can change in arbitrary ways with voltage changes. In addition, signal molecules diffusing out of the dendritic spine (or pumped by its contraction) can interact with these ion channels, changing the way they react to voltage waves. This means that the immediate history of any synapse can have a non-linear effect on how all signals further from the soma along this branch of the dendritic arbor are passed along towards the soma.
Bottom line, the neuron is potentially capable of incredible intelligence. Not only that, but the body contains literally thousands of different types of neurons, each of which can potentially perform a different calculation at each type of synapse it contains. And some nerve cells, such as purkinje cells can have hundreds of thousands of these synapses on a single cell.
Gradients as Calculators2
A gradient of activated enzyme (or even signaling molecule/ion) can act almost as a calculator itself. Such gradients can be created by activating the enzyme at one point and deactivating it somewhere else. The result it that there is a flow of each state of the enzyme from where it's created to where it's converted to another state, and that flow is driven by a concentration gradient. Any other signal gradient (such as from receptors located on the plasma membrane) can interact with it to create balance points within the cell where the outcome of a specific calculation reaches zero. Depending on what other enzymes are located at that balance point, the calculation can result in different actions, translating an analog input into a discrete (digital) output.
When gradients is maintained within the cell, enzymes located on organelles and cellular motors moving along the cytoskeleton can use their location within the cell as input to a calculation that tells them what to do (e.g. where to go) next.
An organelle, or a large enough raft or other structure, can even have several sensing enzymes located far enough apart from one another to determine the direction of the gradient, as well as the strength. This can be used to help decide which of several skeletal components to move along, and which direction to move in, when migrating around the cell.
Long Distance Signaling19, 20, 34
We've seen that many calculations within the Eukaryotic cell are localized (it's somewhat different for prokaryotes). Indeed, the rate of diffusion is so slow within the cytoplasm that high-speed calculations need something else to move them around. We saw that electrical signals can be used (e.g. in nerve and muscle cells), but they have limited capacity to carry information. Other methods that have been proposed are moving vesicles that carry information gleaned from one location to another where it's input to local calculations, and positive feedback cascades which can maintain a very high local concentration gradient at the advancing wave of feedback and thus produce a much higher speed.
Methods I haven't seen proposed, but that are at least possibly feasible, would be moving packages of enzymes that are not membrane enclosed but can hold their state between input and output locations, and localized electrical waves along something like T-tubules within the cell. The latter would be of little use to cells within the body of a multi-celled animal, but single celled creatures might have use for them.
"whew"
This is the last of the series, although there's much more that could be written about "intelligence" in the cell. The most important, how it evolves, requires discussions of complexity theory and is simply too much to include in this series.
Footnotes. I just assembled these and numbered them, so there's no correspondence to the order they're first refereneced in the text. Use the back button if you clicked on a note number above. Not all of the links here are called out in the text.
1. Cellular and subcellular localization of PDE10A, a striatum-enriched phosphodiesterase
2. Cell-signalling dynamics in time and space
3. Phosphoinositide-metabolizing enzymes at the interface between membrane traffic and cell signalling
4. Membrane lipids: where they are and how they behave
5. The function of tight junctions in maintaining differences in lipid composition between the apical and the basolateral cell surface domains of MDCK cells
6. Complex N-Glycan Number and Degree of Branching Cooperate to Regulate Cell Proliferation and Differentiation
7. The vertebrate primary cilium is a sensory organelle
8. Vertebrate Smoothened functions at the primary cilium
9. A Core Complex of BBS Proteins Cooperates with the GTPase Rab8 to Promote Ciliary Membrane Biogenesis
10. Comparative Genomics Identifies a Flagellar and Basal Body Proteome that Includes the BBS5 Human Disease Gene
11. The Chlamydomonas genome reveals the evolution of key animal and plant functions
12. The Mother Centriole Plays an Instructive Role in Defining Cell Geometry
13. Cloning and Characterization of a Novel Human Phosphodiesterase That Hydrolyzes Both cAMP and cGMP
14. Distinct Roles of PDE4 and PDE10A in the Regulation of cAMP/PKA Signaling in the Striatum
15. Diffusion control of protein phosphorylation in signal transduction pathways
16. Signaling cascades as cellular devices for spatial computations
17. Enzyme Localization Can Drastically Affect Signal Amplification in Signal Transduction Pathways
18. Intrinsic Fluctuations, Robustness, and Tunability in Signaling Cycles
19. Four-dimensional organization of protein kinase signaling cascades: the roles of diffusion, endocytosis and molecular motors
20. Modeling the signaling endosome hypothesis: Why a drive to the nucleus is better than a (random) walk
21. Calcium Microdomains in Aspiny Dendrites
22. Signaling in Small Subcellular Volumes. I. Stochastic and Diffusion Effects on Individual Pathways
23. Why the Phosphotransferase System of Escherichia coli Escapes Diffusion Limitation
24. Calcium dynamics in dendritic spines, modeling and experiments
25. Organelles and trafficking machinery for postsynaptic plasticity
26. Modeling calcium dynamics in dendritic spines
27. Calcium Signaling
28. Synaptotagmin arrests the SNARE complex before triggering fast, efficient membrane fusion in response to Ca2+
29. Phosphatidylinositol 4,5-bisphosphate regulates SNARE-dependent membrane fusion
30. Intracellular Trafficking in the Trypanosomatids
31. Intracellular Membrane Transport Systems in Trypanosoma brucei
32. Resurgence of Cardiac T-Tubule Research
33. t-tubules and sarcoplasmic reticulum function in cardiac ventricular myocytes
34. Perinuclear, perigranular and sub-plasmalemmal mitochondria have distinct functions in the regulation of cellular calcium transport
35. Mitochondrial Regulation of Intracellular Ca2+ Signaling: More Than Just Simple Ca2+ Buffers
36. Dynamic molecular confinement in the plasma membrane by microdomains and the cytoskeleton meshwork
37. Mitochondrial-Nuclear Communications
38. Mitochondria: More Than Just a Powerhouse
39. Mitochondria: The Hub of Cellular Ca2+ Signaling
40. Lipid Binding Domains: More than Simple Lipid Effectors
41. Membrane Tubulation by Lattices of Amphiphysin BAR Domains
42. Two distinct but interchangeable mechanisms for flipping of lipid-linked oligosaccharides

Hello,
ReplyDeleteI was very encouraged to find this site. I wanted to thank you for this special read. I can see that you are putting a lots of efforts into your blog. Keep posting the good work.