I'm not going to argue about any of these papers, since two I don't really care about and the third is behind a paywall. However, it brings up an important subject I've discussed before, and probably will again:
The obvious problem with [using cladistics based on parsimony to build a hierarchical classification of species based on phylogeny or evolutionary ancestry] is that our understanding of past evolution is historical in nature, that is we are attempting to determine which of a semi-infinite number of possible evolutionary paths was actually followed in any evolutionary progress. Our estimation of relative probabilities is retrospective, and says little about which path was actually followed.This is important when it comes to paraphyly. According to Wiki, "In phylogenetics, a group of organisms is said to be paraphyletic if the group contains its most recent common ancestor but does not contain all the descendants of that ancestor." The most commonly used example is the "reptiles", a group that traditionally contained all the amniotes (tetrapods that lay air-tolerant eggs or are descended from them) except for birds and mammals.
Consider the following scenario: Species "A" (disregarding problems with the definition of "species") is related to species "B", and "C", but we don't know which it's more closely related to. A cladistic analysis can list a whole bunch of features, some observable in the phenotype, others involving DNA differences such as the deletion or addition of a chunk of DNA, or a point mutation of one base into another, building a table of which species posses which values for which feature. By using several (presumably) less related species as "outgroups" to help determine the ancestral state of each feature, it's possible to build a series of "trees" based on different scenarios for changes to each feature during the evolution leading from the presumed common ancestor to the three modern species.
Parsimony enters in here as the tree with the least number of changes is considered "best", as in most likely. But is it? Suppose the lowest number of changes needed to evolve our three modern species ("A", "B", and "C") is 15. However, there is only one tree involving 15 changes but there are three involving 16, five involving 17, and twelve involving 18 changes. What is the most likely number of changes?
To answer that question, we need to know the relative probability of a change occurring. If such changes are very unlikely, then the tree with 15 changes may be most likely. If changes are fairly likely, then while the tree with 15 changes may be more probable than any single tree with 16, 17, or 18, the probability that the actual (historical) evolution had some other number of changes (probably 18, considering the number of possible paths) becomes higher than that for 15 changes.
This is not a trivial question, when we go beyond static cladistics to ask how each species came to evolve. In addition to mutations that cause the changes, we also need to consider the selective pressure that went into fixing those changes into the lineage that evolved into one of our modern species. It may be that the likelihood that the most parsimonious path was followed is low compared to one in which an ancestor of two modern species followed some sort of zig-zag path, adding changes that were later reversed, adding the same change more than once (especially for morphological features that were directly driven by selection, etc.)
Now, what Williams and Ebach (the writers of Systematics and Biogeography) are unhappy about it the way traditional taxonomists are attempting to defend older groups that are paraphyletic: ...
Brummitt's Evolution in taxonomic perspective consists of several abuses of paraphyly:This may be true of Brummitt, I haven't read the paper. But based on the quote here, I can see a perfectly reasonable meaning: the word "evolution" here actually is a short-hand for "evolutionary process", which in turn is concerned with much more than simple ancestry: the how and why of the evolution."... emphasis has been increasingly placed on the need for a classification which recognises evolution" (Brummitt, 2008:1050).This is what monophyly is all about. Once you discover monophyly you have discovered evolution within your group. Unfortunately, this is contrary to Brummitt (and his followers) who believe that their taxonomies alone (without any need for testing, it seems) are evolutionary. This runs counter to empiricism in science, which makes hypotheses of relationships (e.g., taxonomies) and uses cladistics to test them. It seems that Brummitt's taxonomy is the yardstick that cladistics has to abide. If it doesn't (i.e., the taxon is paraphyletic) cladistics is wrong, not Brummitt. Therefore paraphyly is evidence and empiricism politely excused
Reptiles: an Example
Let's go back to the reptiles: "cold-blooded", sprawled stanced, relatively small-brained (also simple brained). Both the birds and the mammals depart from this structure by being "warm-blooded", upright stanced, and possessing much more sophisticated brains. Although this simple picture becomes much more complex when we actually study the lineages leading to the mammals and birds (the latter including most of the more impressive dinosaurs), from the point of view of actually trying to recreate the evolutionary track followed, the "mammal-like reptiles" and the archosaurs followed a divergent path, evolving away from the basic characteristics of the presumed ancestral lineage.
Thus, it makes sense from the point of view of looking at the evolutionary process to create a "class" reptilia which performs a series of adaptive radiations based around the original evolutionary inovations of the first amniote, while allowing lineages such as the archosaurs and therapsids to "leave" that class and found a new one based on a new wave of innovations.
It's important to recognize the limitations of such paraphyletic classifications. In order to be useful in modeling the evolutionary process, they must retain the primary advantageous innovations of the original ancestor, without adding any new innovations that set of a new round of adaptive radiation. Creating a group called "reptiles" that includes the early synapsids but excludes the dinosaurs and birds would be of little use, since at least some of the innovations that set the mammals apart are also present in many dinosaurs and the birds. A "class reptilia", to be useful in modeling evolution, would have to remain predominantly cold-blooded, sprawled stanced, egg-laying, and simple-brained.
OTOH, the occasional viviparous (or oviviparous) snake or lizard can be retained unless it stands at the base of a major adaptive radiation. The turtles would be a borderline case (IMO), if they had had a much larger adaptive radiation they would reasonably count as another departure.
An Adventure in Morphospace
I'm going to get a bit more technical here, and consider what adaptive radiation does, and how I think we should distinguish between a lineage that may be removed from its parent lineage to leave a useful paraphyletic classification. It begins with morphospace.
Wiki doesn't provide a definition of morphospace, so, with the aid of a page I found through Google, let me expound.
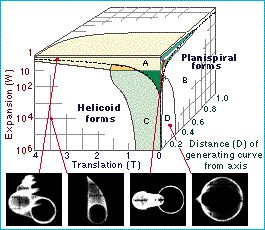
Figure 1: Morphospace schematic. Click on image to see original caption. (From the website of Mark Ridley's Evolution)
Basically, you have to imagine an n-dimensional space with each of the n axes representing the extent of one characteristic of the morphology. The location in that space represents the specific morphology. An adaptive radiation spreads out in this space, along one or several dimensions.
A key innovation, OTOH, involves a movement along dimensions not really involved in previous adaptive radiations. Thus, early "reptiles" spread out in various ways based on their "lizard-like" shape, their "cold blood", and minor modifications of their body plans and egg styles, while the lineages leading to birds and mammals innovated with upright stance, "warm blood", milk, feathers capable of flight, and so on. These key innovations opened up a new area of morphospace for adaptive radiation, meaning that a qualitatively different evolutionary process was followed.
This, in my view, represents a good justification for allowing paraphyletic classes such as "reptiles": they help in modeling the evolutionary process, because the divergence involved in creating the lineages that "leave" the class are qualitatively different from those involved in the adaptive radiations within it.
Obviously, there are some semantic boundary problems here. In some ways, the differences between the types of evolution involved in simple adaptive radiation and the development of the archosaurs and mammals can be seen as differences in degree rather than kind. However, we can judge by the results, and also by the likelihood of the development. If mutations leading to a particular development were happening all the time, there would have been many instances of it. OTOH, if we're talking about the kind of mutation whose probability of occurrence is (on average) once in every 100 million years for a large population, then we can probably allow such mutations to count as "key" innovations when they lead to a major adaptive radiation.
Cladistic Reconciliation
How can this be reconciled with the more extreme cladistic demands? Well, we need to start by realizing that these "grade-based" paraphyletic classifications are only useful tools for modeling evolutionary processes, they aren't for resolving issues of "genetic relatedness". We need to carefully distinguish between genetic relatedness and the morpho-ecological "relationships" that tie together creatures with similar morphology and ecological niches. When we are discussing ancestry and genetic relatedness we need to concentrate on cladistic analyses, allowing some modification based on specific theoretical evolutionary scenarios.
Only when we're trying to build those scenarios, especially when the cladistic issues cannot be resolved (as is the case, for instance, with the earliest animals), should we allow discussion of paraphyletic classifications such as "reptiles" or "sponges".

No comments:
Post a Comment